Membrane Protein Conference on September 15, 2026
Our next Membrane Protein Conference will be held on Tuesday Sept 15, 2026 from 2-5pm CET (8-11 am ET) and will feature a series of 20 minute talks covering the latest findings about membrane structure and function, lipid-protein interactions as well as advances for purification and resolution of native membrane assemblies.
We are seeking to explore and disseminate new concepts and tools in the field of membrane protein biology. Free student tickets are available, and registration is 20 or 30 euros for academics or industry to help cover our costs as an independent organization. Dip in and out of this zoom meeting as needed and receive the recordings afterwards.
Speakers at the MPC on September 15, 2026 include:
 Alexej Kedrov, Associate Professor, University of Düsseldorf on the substrate-induced assembly and functional mechanism of the bacterial membrane protein insertase SecYEG-YidC using nanodiscs and cryo-EM, which has just been accepted in EMBO J.
Alexej Kedrov, Associate Professor, University of Düsseldorf on the substrate-induced assembly and functional mechanism of the bacterial membrane protein insertase SecYEG-YidC using nanodiscs and cryo-EM, which has just been accepted in EMBO J.
 Surbhi Dhingra, Postdoctoral Fellow with Kenton Swartz, National Institute of Neurological Disorders and Stroke, NIH, who recently published Mechanisms of ligand recognition and channel opening for P2X2 receptors in lipid nanodiscs in Science Advances.
Surbhi Dhingra, Postdoctoral Fellow with Kenton Swartz, National Institute of Neurological Disorders and Stroke, NIH, who recently published Mechanisms of ligand recognition and channel opening for P2X2 receptors in lipid nanodiscs in Science Advances.
 George Hedger, Senior Postdoc with Sarah Rouse, Department of Life Sciences, Imperial College London, on native mass spectrometry, multiscale simulations, machine learning, and fluorescence microscopy of GPCR targets; recently published Ligand-like lipid interactions with membrane proteins: Simulations and machine learning in COSB.
George Hedger, Senior Postdoc with Sarah Rouse, Department of Life Sciences, Imperial College London, on native mass spectrometry, multiscale simulations, machine learning, and fluorescence microscopy of GPCR targets; recently published Ligand-like lipid interactions with membrane proteins: Simulations and machine learning in COSB.
 Moitrayee Bhattacharyya, Assistant Professor of Pharmacology, Department of Pharmacology, School of Medicine, Yale University, on Single molecule microscopy and cryo-EM with native nanodiscs in understanding neurodegeneration and pain.
Moitrayee Bhattacharyya, Assistant Professor of Pharmacology, Department of Pharmacology, School of Medicine, Yale University, on Single molecule microscopy and cryo-EM with native nanodiscs in understanding neurodegeneration and pain.
 Shiv K. Sah Teli, Postdoctoral Researcher with Juha Huiskonen, Laboratory of Structural Biology, University of Helsinki, who published Gating Mechanism of the Human Connexin 45 Gap Junction in bioRxiv.
Shiv K. Sah Teli, Postdoctoral Researcher with Juha Huiskonen, Laboratory of Structural Biology, University of Helsinki, who published Gating Mechanism of the Human Connexin 45 Gap Junction in bioRxiv.
 Salma Dienta Salsabila, PhD Student with Jungwook Kim, Gwangju Institute of Science and Technology, who published Structural mechanism of membrane-associated cytidine diphosphate diacylglycerol diphosphatase in Escherichia coli. in Int J Biol Macromol.
Salma Dienta Salsabila, PhD Student with Jungwook Kim, Gwangju Institute of Science and Technology, who published Structural mechanism of membrane-associated cytidine diphosphate diacylglycerol diphosphatase in Escherichia coli. in Int J Biol Macromol.
MPC on May 12, 2026 Video, Attendees and Program:
13:50 CET Informal opening with introductions and sharing of tips & tricks
Session 1 chaired by Gestél Kuyler, CEO, Nanosene
 14:00 Christos Pliotas, Reader in Structural Biological EPR Spectroscopy, Division of Molecular & Cellular Function, School of Biological Sciences, University of Manchester, recently published Swapped and non-swapped TRAAK states co-exist in membranes at a ratio influenced by temperature in Nat. Commun.
14:00 Christos Pliotas, Reader in Structural Biological EPR Spectroscopy, Division of Molecular & Cellular Function, School of Biological Sciences, University of Manchester, recently published Swapped and non-swapped TRAAK states co-exist in membranes at a ratio influenced by temperature in Nat. Commun.
 14:20 Emmanuelle Quemin, Group Leader, I2BC, CNRS will speak on probing virus-host interactions using cellular cryo electron tomography, and published Subcellular reorganization upon phage infection reveals stepwise assembly of viral particles from membrane-associated precursors in Nat Commun.
14:20 Emmanuelle Quemin, Group Leader, I2BC, CNRS will speak on probing virus-host interactions using cellular cryo electron tomography, and published Subcellular reorganization upon phage infection reveals stepwise assembly of viral particles from membrane-associated precursors in Nat Commun.
 14:40 Alexandra Steinhauser, PhD student, Faculty of Science and Engineering, U Groningen, who recently published Home is where the lipids are: a comparison of MSP and DDDG nanodiscs for membrane protein research in Soft Matter.
14:40 Alexandra Steinhauser, PhD student, Faculty of Science and Engineering, U Groningen, who recently published Home is where the lipids are: a comparison of MSP and DDDG nanodiscs for membrane protein research in Soft Matter.
15:00 Break with Q&A
Session 2 chaired by Evelyn Okorafor, PhD Student, Miami University
 15:10 Adel Hussein, Postdoctoral Fellow, Department of Biochemistry and Molecular Pharmacology, New York University with David Stokes, will present Lipids are essential for potassium transport by KdpFABC from E. coli, which is on bioRxiv.
15:10 Adel Hussein, Postdoctoral Fellow, Department of Biochemistry and Molecular Pharmacology, New York University with David Stokes, will present Lipids are essential for potassium transport by KdpFABC from E. coli, which is on bioRxiv.
 15:30 Yasmin Aydin, Post-doctoral Researcher at Indiana University School of Medicine with Qiuyan Chen will present Lipid Zipcodes and Phosphorylation Barcodes Cooperatively Control Arrestin Activation as is also discussed in her bioRxiv article.
15:30 Yasmin Aydin, Post-doctoral Researcher at Indiana University School of Medicine with Qiuyan Chen will present Lipid Zipcodes and Phosphorylation Barcodes Cooperatively Control Arrestin Activation as is also discussed in her bioRxiv article.
 15:50 Eugene Agyemang PhD Student, Biochemistry & Cellular and Molecular Biology, U Tennessee, Knoxville with Rajan Lamichhane, who recently published on Activation dynamics of a water-soluble human mu-opioid receptor in JBC.
15:50 Eugene Agyemang PhD Student, Biochemistry & Cellular and Molecular Biology, U Tennessee, Knoxville with Rajan Lamichhane, who recently published on Activation dynamics of a water-soluble human mu-opioid receptor in JBC.
Session 3 chaired by Youzhong Guo, Associate Professor, VCU
16:10 Break with Q&A
 16:20 Amedee des Georges, Associate Professor, Molecular Pathobiology, NYU College of Dentistry and Investigator in the NYU Pain Research Center, will present on the mechanism of beta-arrestin 1 mediated Src activation via Src SH3 domain revealed by c-EM.
16:20 Amedee des Georges, Associate Professor, Molecular Pathobiology, NYU College of Dentistry and Investigator in the NYU Pain Research Center, will present on the mechanism of beta-arrestin 1 mediated Src activation via Src SH3 domain revealed by c-EM.
 16:40 Cristina Santarossa, Postdoctoral Fellow in Gira Bhabha's lab at Johns Hopkins University, who recently published LetA defines a structurally distinct transporter family in Nature.
16:40 Cristina Santarossa, Postdoctoral Fellow in Gira Bhabha's lab at Johns Hopkins University, who recently published LetA defines a structurally distinct transporter family in Nature.
17:00 Discussion and close
Fill out our short Conference Evaluation Form, we value your feedback.
Speaker Awards
Sponsored by Nanosene. Our previous winners:
- Alyssa Ward, PhD Student, U Tennessee Knoxville
- Kirill Nadezhdin, Columbia University Irving Medical Center
- Rhythm Shukla, University of Utrecht
- David Glück, Karl-Franzens-Universität Graz
- Johanna Syrjanen, Cold Spring Harbor Laboratory
- Gestél Kuyler, Stellenbosch & Coventry Universities
- Elaine Bernard, graduate student, Stellenbosch U
- Evelyn Okorafor, graduate student, Miami U
- Ruby Huynh, undergraduate student, Arizona State U
- Arba Zhao, high school student at Webb High School
- Elissa Moller, PhD student, NICHD
- Claire Coupland, Postdoc, UToronto
- Frank Tucci, PhD student at Northwestern U
- Surabhi Kokane, PhD student, Stockholm U
- Mackenzie Thompson, Postdoctoral Researcher, UC Berkeley
Background on SMALP
The SMALP network is an open forum initiated by the international research community. We are developing and using amphipathic copolymers as alternatives to conventional detergents, which tend to dissociate, destabilize and deactivate native membrane proteins.
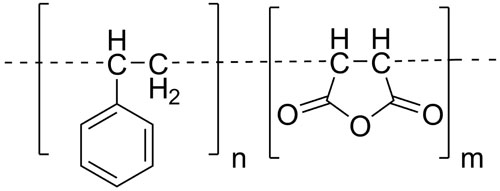
A variety of amphipathic polymers such as styrene maleic acid have been found to directly and spontaneously solubilize proteins and bound lipids from membranes when activated. This provides opportunities for understanding and exploiting the structure and function of native membrane protein complexes.
As SMALP technology is relatively new, the participation of the wider community and sharing of new ideas and practices amongst researchers is the best way to ensure continued rapid development of the field.
The SMALP network offers polymers, methods and meetings to support the growing community of scientists and companies interested in the preparation of membrane nanodiscs formed by polymers including styrene maleic acid for biological research and drug discovery applications.
Register for a day of insightful talks from experts in the structural biology and functions of membrane proteins and nanodisc development, discussions with academic and industry leaders, and enjoy breakfast, lunch and coffee breaks on us.
These events are offered to bring together academic and industry experts, postdocs and students who are interested in membrane protein structural biology, lipidomics, proteomics, target discovery and biophysical analysis of bilayers and amphipathic polymers.
We welcome researchers from across academia and industry to join us in lively discussions and debate about the latest advances in native nanodiscs, membrane proteins, polymer chemistry, lipid biophysics, structural biology, and applications including drug discovery and delivery.

